Content :
The API Biochemical Gallery is a testing system commonly used in microbiology to aid in the identification of bacteria. In this article, we will explore the principle, usage, and interpretation of the API system strip.
◉ Introduction
API test strip (analytical profile index) is a miniaturized and standardized gallery of biochemical tests, usable with complete identification databases, the best known of which is the api 20E (20 characters for Enterobacteriaceae).
According to BIOMERIEUX, 10 API® strips are used every minute worldwide, they are also used to assess the performance of other identification products (reference technique)
Initial identification of bacteria via methods such as Gram staining or selective or differential culture media is essential before choosing the appropriate API gallery.
Similar systems are offered by other companies, such as the Liofilchem EnteroPluri-Test.
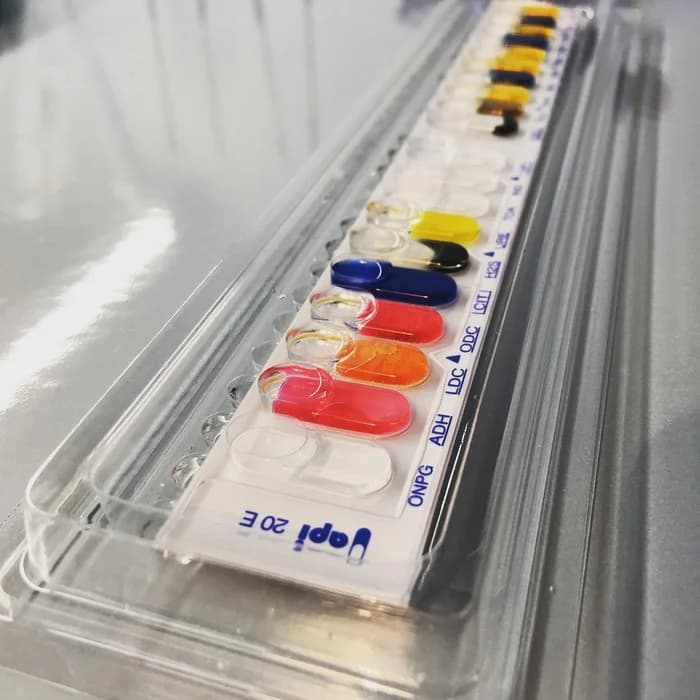

◉ API 20E identification gallery
The API 20E includes a plastic strip containing 20 mini test wells. Each well contains dehydrated media with chemically defined compositions to generally detect enzymatic activity, primarily related to carbohydrate fermentation or protein and amino acid catabolism by the inoculated organisms.
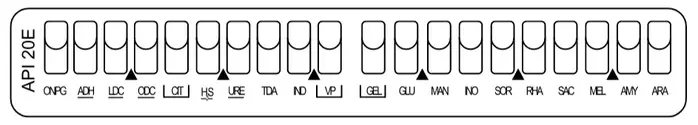
◉ The different tests in API 20E
1- ONPG test : The ONPG test is used to detect the enzyme β-galactosidase
2- ADH : Decarboxylation of the amino acid arginine by arginine dihydrolase
3- LDC: Decarboxylation of the amino acid lysine by lysine decarboxylase
4- ODC: Decarboxylation of the amino acid ornithine by ornithine decarboxylase
5- CIT : Use of citrate as sole carbon source
6- H2S : Hydrogen sulfide production
7- URE : Urease enzyme test
8- TDA (Tryptophan deaminase) : Detection of the tryptophan deaminase enzyme (detected by adding Ferric chloride).
9- IND : Indole test : production of indole from tryptophan by the enzyme tryptophanase (Indole is detected by adding Kovac's reagent).
10- VP : The Voges-Proskauer test for the detection of acetoin (acetylmethylcarbinol) produced by fermentation of glucose by bacteria using the butylene glycol pathway
11- GEL : Production test of the enzyme gelatinase which liquefies gelatin
12- GLU : Glucose fermentation (hexose sugar)
13- MAN : Fermentation of mannose (sugar hexose)
14- INO : Fermentation of inositol (cyclic polyalcohol)
15- SOR : Fermentation of sorbitol (sugar alcohol)
16- RHA : Rhamnose fermentation (methyl pentose sugar)
17- SAC : Fermentation of sucrose (disaccharide)
18- MEL : Fermentation of melibiose (disaccharide)
19- AMY : Fermentation of amygdalin (glycoside)
20- ARA Fermentation of arabinose (pentose sugar)
A bacterial suspension is used to rehydrate each well, and the strips are then incubated. During this incubation, the metabolic activities of the bacteria cause color changes, which appear spontaneously or are revealed by the addition of reagents.
◉ Preperation of the API gallery
❶ Take a single isolated colony (from a pure culture) and make a bacterial suspension in sterile distilled water (0.5 McFarland).
❷ Combine the bottom and lid of an incubation box and distribute water in the cells to create a humid atmosphere. Then place the gallery in the incubation box under sterile conditions
❸ Take a Pasteur pipette and fill these compartments with the bacterial suspension as follows (photo) :
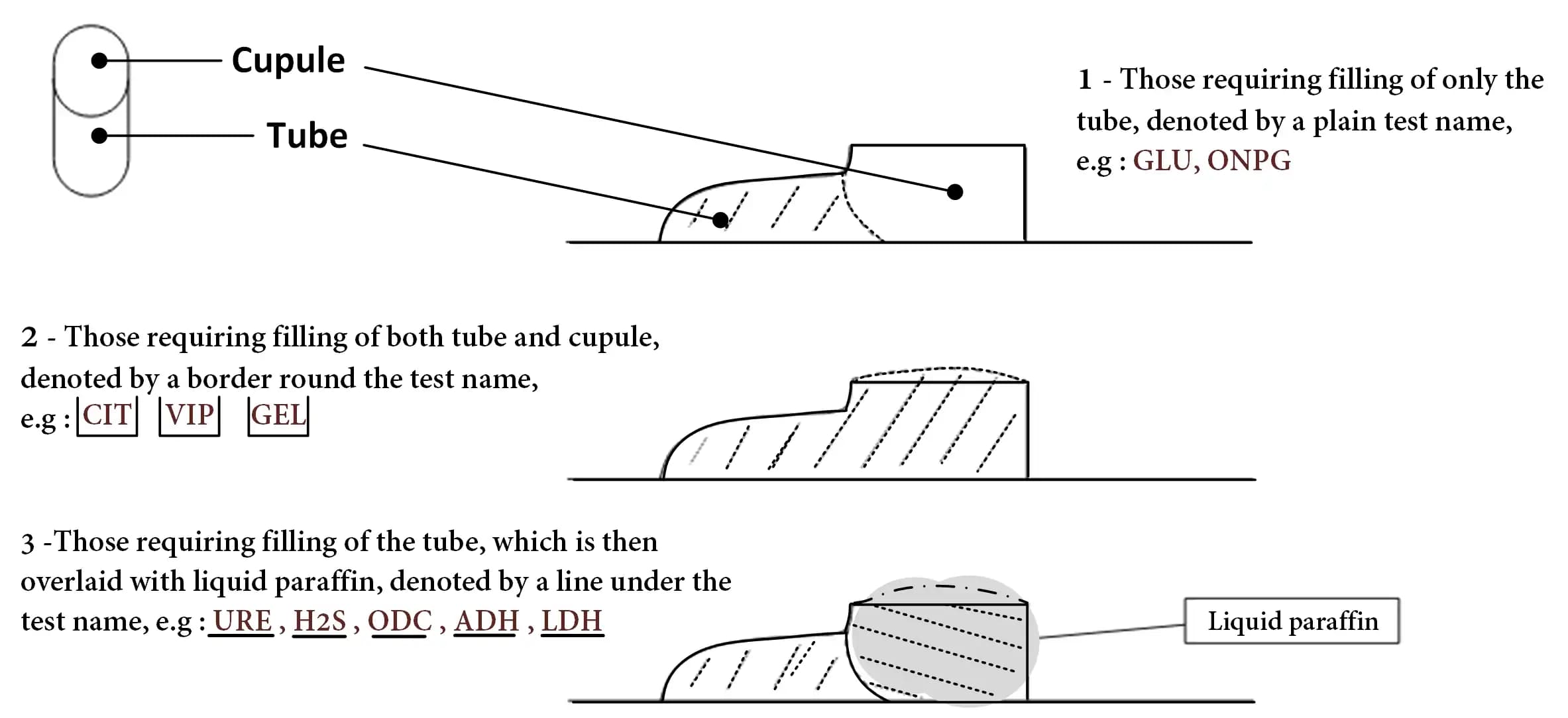
❹ Mark the tray with the identification number (Patient ID or Organization ID), date and your initials.
❺ Incubate the tray at 37 o C for 18-24 hours.
❻ After incubation add reagents as follows :
| Well | Reagent |
|---|---|
| TDA | A drop of TDA reagent |
| IND | A drop of James' reagent or kovacs |
| VP | A drop of VP1 then VP2 reagent |
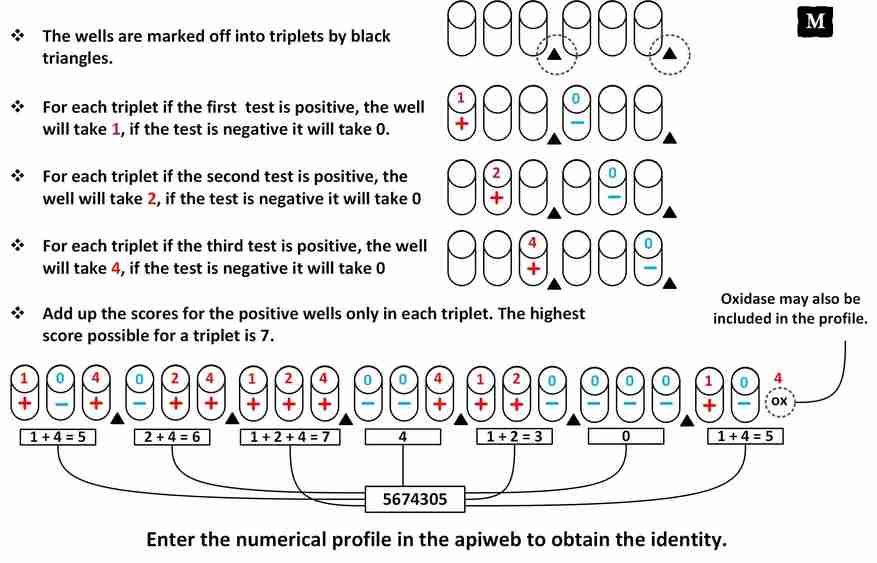
◉ Results, reading and interpretation of the API 20E
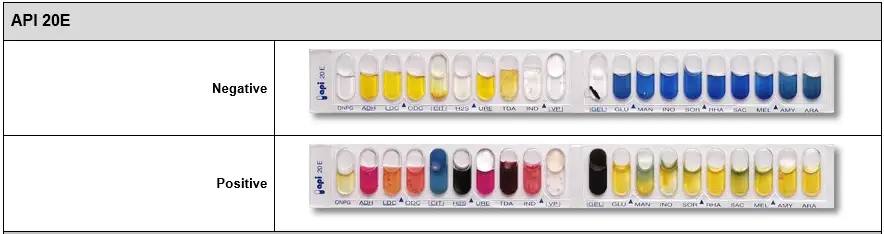
The reading of these reactions (positive or negative) is done according to the variations of colors
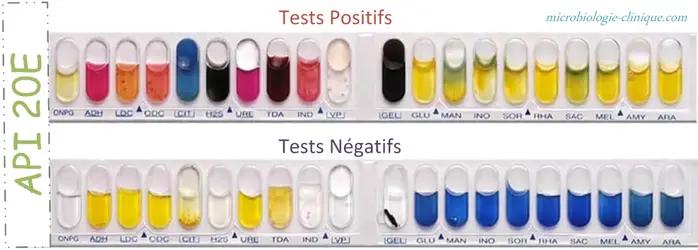

API 20E positive vs negative
- You can use a free online tool to interpret your API test, such as UPBM (note that the database may not be up to date).
- You can also enter your results into the Excel spreadsheet Excel spreadsheet
- Another option is to use apiwebTM biomerieux
◉ Download Protocols from API Galleries
🏾 We have collected the main manuals to carry out your biochemical identification API (API 20e, API NE, API STREPT, API STAPH, API CANDIDA...)
:| Galleries | Protocols |
|---|---|
| API 20E pdf | Download |
| API NE pdf | Download |
| API STREPT pdf | Download |
| API STAPH pdf | Download |
| API LISTERIA pdf | Download |
| API CANDIDA pdf | Download |
| API CH50 pdf | Download |
| API 20C pdf | Download |
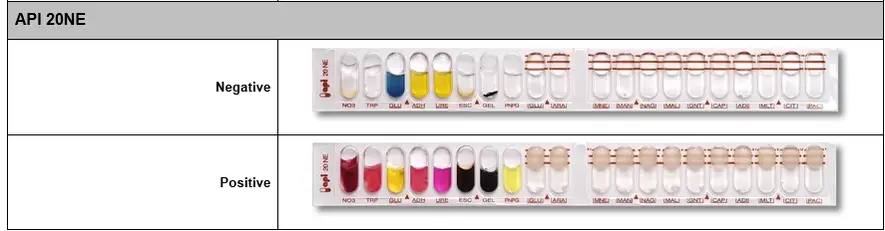
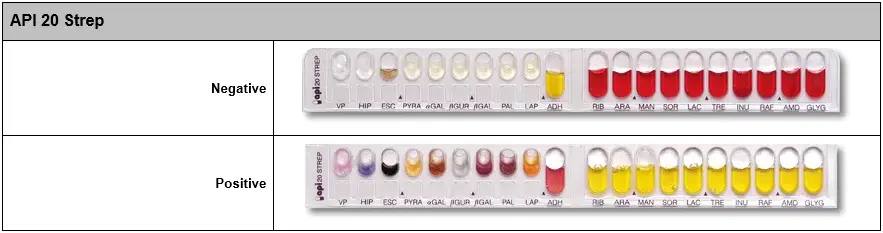
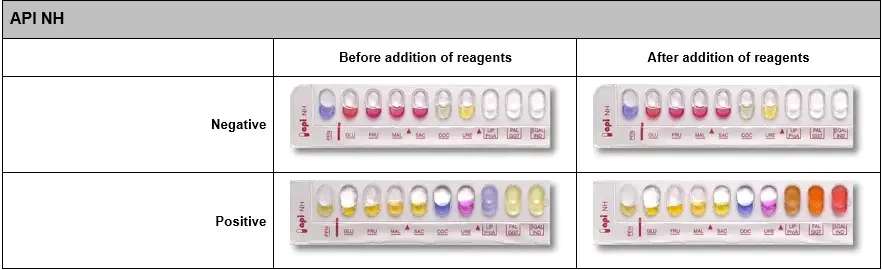
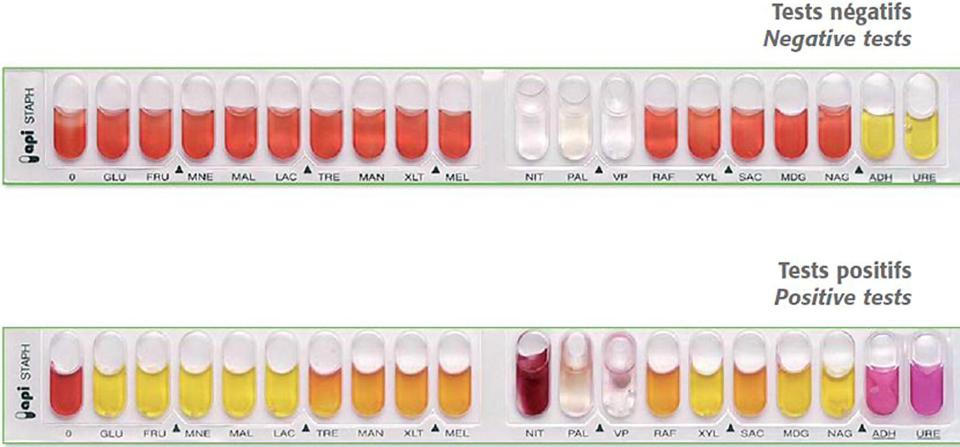
You can have all the information necessary for the handling and interpretation of API 20e, API NE, API STREPT, API STAPH, API CANDIDA... in the following file Download